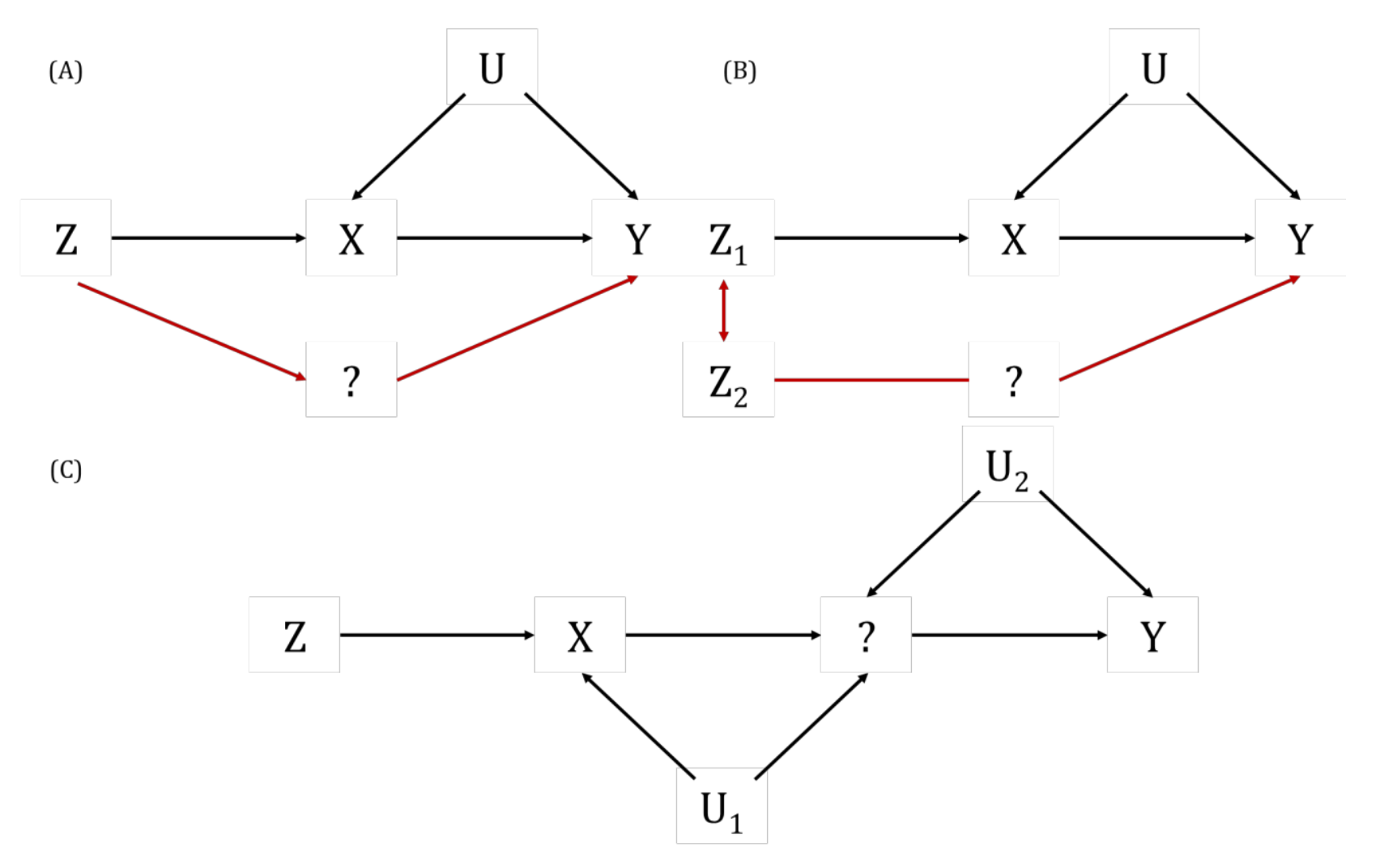
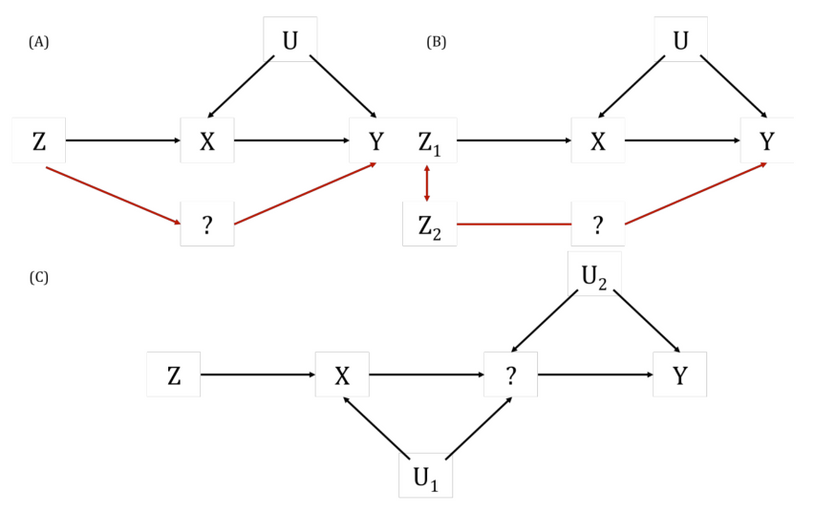
Also known as “spurious” or “false” pleiotropy. This is when a genetic variant affects other traits (which influence the outcome) via its effect on the exposure.
This is the very phenomenon that MR seeks to detect (i.e., the chain of traits to the outcome is what gives rise to an unbiased non-null causal effect estimate).
References
- Davey Smith G, Hemani G. Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum Mol Genet 2014; 23: R89-R98.
- Zheng J, Baird D, Borges MC, et al. Recent Developments in Mendelian Randomization Studies. Curr Epidemiol Rep 2017; 4: 330-345.
Other terms in 'Sources of bias and limitations in MR':
- Assortative mating
- Canalization
- Collider
- Collider bias
- Conditional F-statistic for multiple exposures
- Confounding
- Exclusion restriction assumption
- F-statistic
- Harmonization (in two-sample MR)
- Homogeneity Assumption
- Horizontal Pleiotropy
- Independence assumption
- INstrument Strength Independent of Direct Effect (InSIDE) assumption
- Intergenerational (or dynastic) effects
- Monotonicity assumption
- MR for testing critical or sensitive periods
- MR for testing developmental origins
- No effect modification assumption
- NO Measurement Error (NOME) assumption
- Non-linear MR
- Non-overlapping samples (in two-sample MR)
- Overfitting
- Pleiotropy
- Population stratification
- R-squared
- Regression dilution bias (attenuation by errors)
- Relevance assumption
- Reverse causality
- Same underlying population (in two-sample MR)
- Statistical power and efficiency
- Weak instrument bias
- Winner's curse